cKBET: assessing goodness of batch effect correction for single-cell RNA-seq
Por um escritor misterioso
Last updated 12 abril 2025

lt;p>Single-cell RNA sequencing reveals the gene structure and gene expression status of a single cell, which can reflect the heterogeneity between cells. However, batch effects caused by non-biological factors may hinder data integration and downstream analysis. Although the batch effect can be evaluated by visualizing the data, which actually is subjective and inaccurate. In this work, we propose a quantitative method cKBET, which considers the batch and cell type information simultaneously. The cKBET method accesses batch effects by comparing the global and local fraction of cells of different batches in different cell types. We verify the performance of our cKBET method on simulated and real biological data sets. The experimental results show that our cKBET method is superior to existing methods in most cases. In general, our cKBET method can detect batch effect with either balanced or unbalanced cell types, and thus evaluate batch correction methods.</p>

CellMixS: quantifying and visualizing batch effects in single-cell RNA-seq data

PDF) A benchmark of batch-effect correction methods for single-cell RNA sequencing data

PDF) BERMUDA: A novel deep transfer learning method for single-cell RNA sequencing batch correction reveals hidden high-resolution cellular subtypes
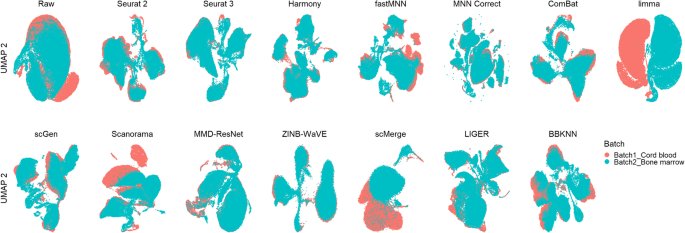
A benchmark of batch-effect correction methods for single-cell RNA sequencing data, Genome Biology

PDF) A geometrical approach based on PCA to benchmark the algorithms of batch effect correction applied to the integration of RNA-Seq data
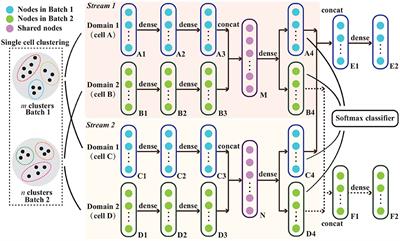
Frontiers CBA: Cluster-Guided Batch Alignment for Single Cell RNA-seq
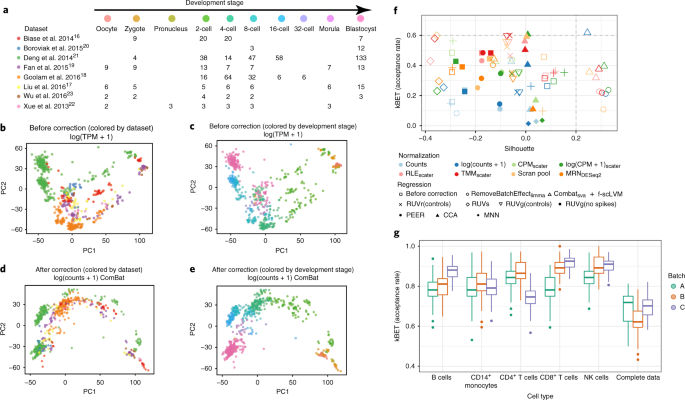
A test metric for assessing single-cell RNA-seq batch correction

Assessment of batch-correction methods for scRNA-seq data with a new test metric
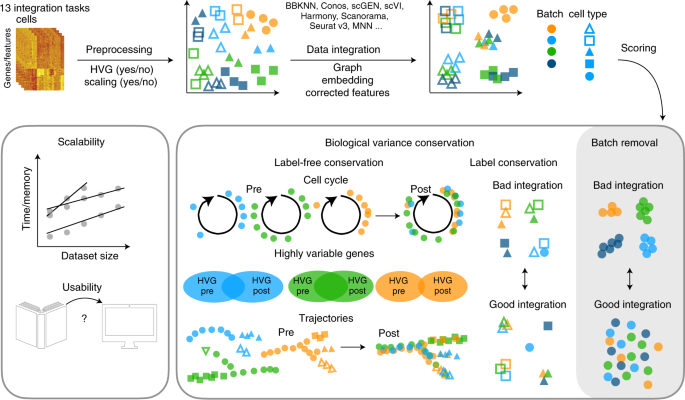
Benchmarking atlas-level data integration in single-cell genomics
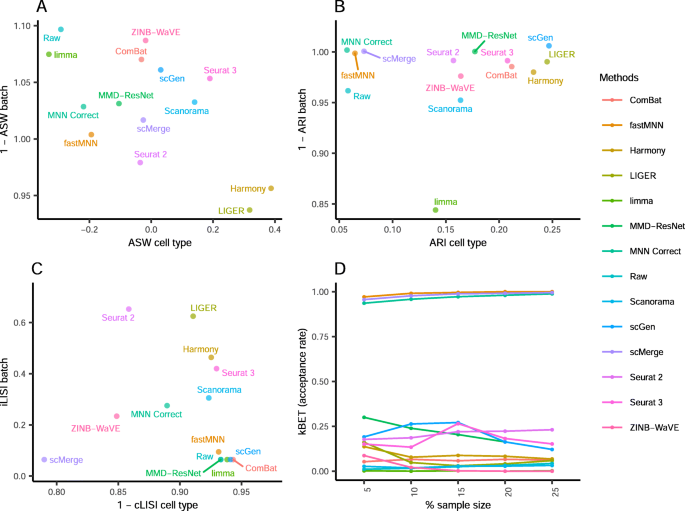
A benchmark of batch-effect correction methods for single-cell RNA sequencing data, Genome Biology
Recomendado para você
-
 CKBET APK for Android Download12 abril 2025
CKBET APK for Android Download12 abril 2025 -
 Ckbet Baixar aplicativo para Android ou iOS 202312 abril 2025
Ckbet Baixar aplicativo para Android ou iOS 202312 abril 2025 -
CKbet - 🦢CKbet🦢 🔥 CKBET Members Committee🔥 Be a CKBET agent, invite your friends to be part of your team and get your commission without even having to bet or play anything!12 abril 2025
-
 Ckbet - Ckbet casino - Casinos Online Famosos no Brasil12 abril 2025
Ckbet - Ckbet casino - Casinos Online Famosos no Brasil12 abril 2025 -
 Browse thousands of 168jogo Com Sacar Ckbet Com95429 images for design inspiration12 abril 2025
Browse thousands of 168jogo Com Sacar Ckbet Com95429 images for design inspiration12 abril 2025 -
 Ckbet – Oferecendo Uma Experiência de Apostas Segura e Conveniente Para Usuários Brasileiros – Portal G3712 abril 2025
Ckbet – Oferecendo Uma Experiência de Apostas Segura e Conveniente Para Usuários Brasileiros – Portal G3712 abril 2025 -
CKbet - Comissão de Membros CKBET Seja um agente da CKBET, convide seus amigos para fazerem parte da sua equipe e obtenha sua comissão sem nem precisar apostar ou jogar nada! Contanto12 abril 2025
-
Ckbet APK (Android App) - Free Download12 abril 2025
-
 ckbet - Seu Portal para Jogos Online Empolgantes.12 abril 2025
ckbet - Seu Portal para Jogos Online Empolgantes.12 abril 2025 -
 CKBET Paga Mesmo? Dá pra ganhar dinheiro?12 abril 2025
CKBET Paga Mesmo? Dá pra ganhar dinheiro?12 abril 2025
você pode gostar
-
 Total Drama Presents:The Ridonculous Race by EerioHydro12345 on DeviantArt12 abril 2025
Total Drama Presents:The Ridonculous Race by EerioHydro12345 on DeviantArt12 abril 2025 -
 Oxford Xadrez (Italiano)12 abril 2025
Oxford Xadrez (Italiano)12 abril 2025 -
 Outra vez arroz, Maria João?12 abril 2025
Outra vez arroz, Maria João?12 abril 2025 -
 Best Buy: Five Nights at Freddy's Security Breach PlayStation 412 abril 2025
Best Buy: Five Nights at Freddy's Security Breach PlayStation 412 abril 2025 -
 Spider-Man: Far From Home Ending Scene12 abril 2025
Spider-Man: Far From Home Ending Scene12 abril 2025 -
 RedEngine Cracked12 abril 2025
RedEngine Cracked12 abril 2025 -
Parabens Julinha 🎂 thanks for being the light in my life , we love you ✨❤️12 abril 2025
-
 The Key West Flagship The Western Union Schooner Tied Up At The12 abril 2025
The Key West Flagship The Western Union Schooner Tied Up At The12 abril 2025 -
 Counter-Strike 2 is dragging down CSGO's reputation12 abril 2025
Counter-Strike 2 is dragging down CSGO's reputation12 abril 2025 -
 Ox, Trade Roblox Adopt Me Items12 abril 2025
Ox, Trade Roblox Adopt Me Items12 abril 2025


